{batsch} provides real-time PCR data sets by Batsch et
al. (2008) in tidy format. There are five data sets bundled with this
package, one for each PCR target:
SLC6A14r for rat SLC6A14SLC22A13h for human SLC22A13EMTp for pig EMTETTch for chicken ETTGAPDHh for human GAPDHEach data set comprises a five-point, four-fold dilution series. For each concentration there are three replicates. Each amplification curve is 45 cycles long.
Install {batsch} from CRAN:
# Install from CRAN
install.packages("batsch")You can instead install the development version of
{batsch} from GitHub:
# install.packages("remotes")
remotes::install_github("ramiromagno/batsch")library(batsch)
SLC6A14r %>%
ggplot(mapping = aes(
x = cycle,
y = fluor,
group = interaction(dilution, replicate),
col = as.factor(dilution)
)) +
geom_line(linewidth = 0.5) +
labs(y = "Raw fluorescence", colour = "Dilution factor", title = "Rat SLC6A14") +
guides(color = guide_legend(override.aes = list(size = 0.5)))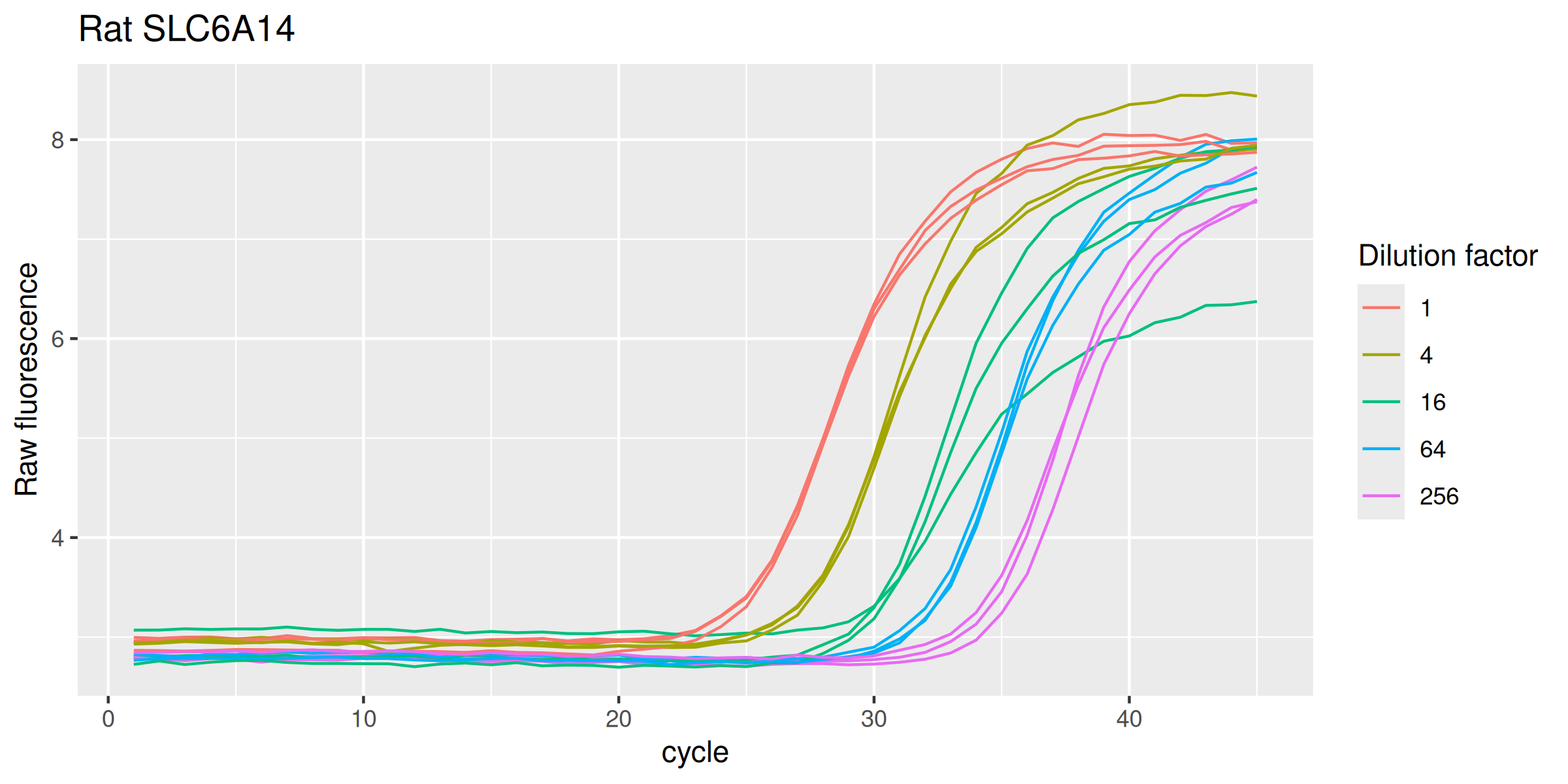
SLC22A13h %>%
ggplot(mapping = aes(
x = cycle,
y = fluor,
group = interaction(dilution, replicate),
col = as.factor(dilution)
)) +
geom_line(linewidth = 0.5) +
labs(y = "Raw fluorescence", colour = "Dilution factor", title = "Human SLC22A13") +
guides(color = guide_legend(override.aes = list(size = 0.5)))
EMTp %>%
ggplot(mapping = aes(
x = cycle,
y = fluor,
group = interaction(dilution, replicate),
col = as.factor(dilution)
)) +
geom_line(linewidth = 0.5) +
labs(y = "Raw fluorescence", colour = "Dilution factor", title = "Pig EMT") +
guides(color = guide_legend(override.aes = list(size = 0.5)))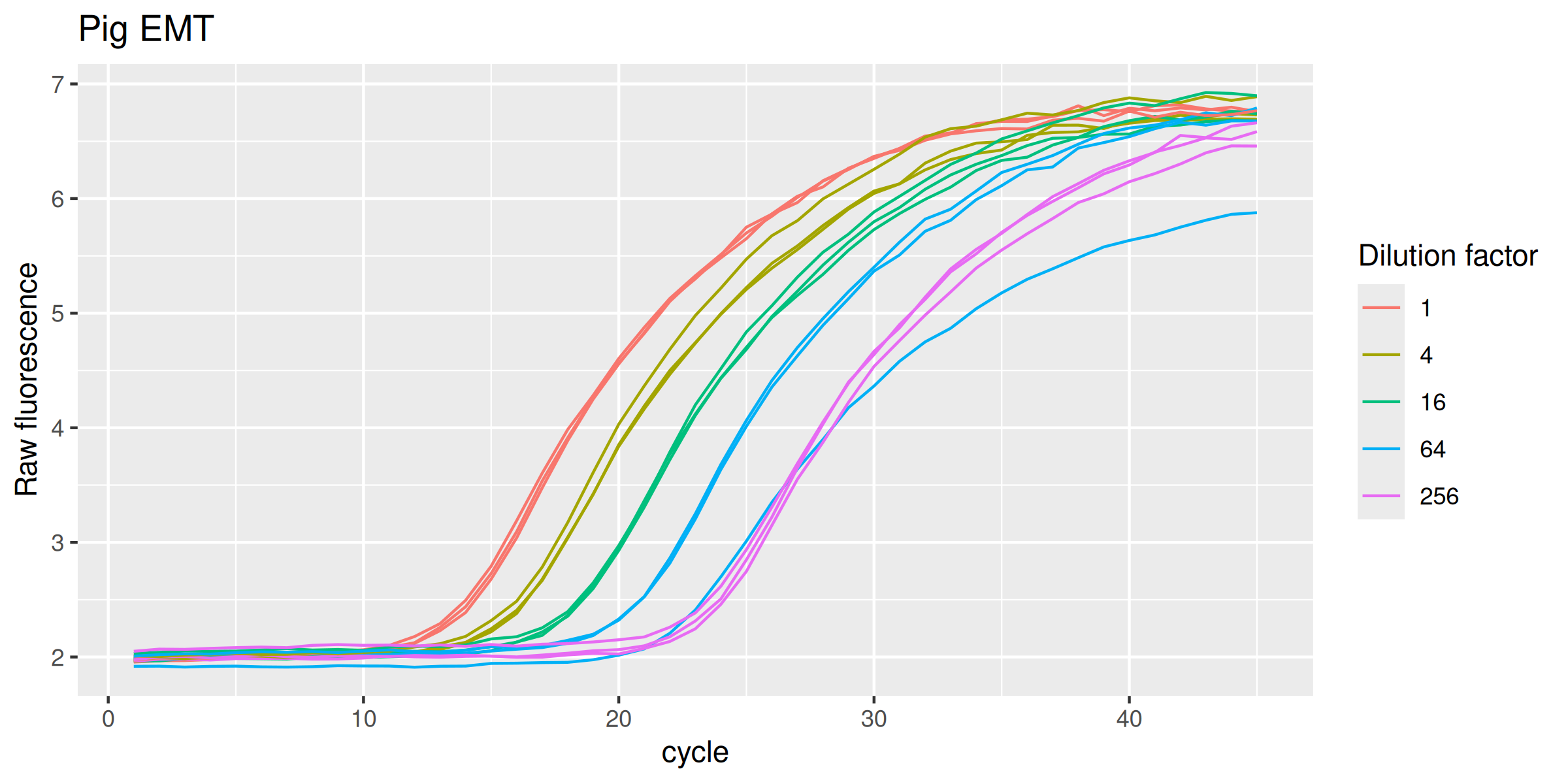
ETTch %>%
ggplot(mapping = aes(
x = cycle,
y = fluor,
group = interaction(dilution, replicate),
col = as.factor(dilution)
)) +
geom_line(linewidth = 0.5) +
labs(y = "Raw fluorescence", colour = "Dilution factor", title = "Chicken ETT") +
guides(color = guide_legend(override.aes = list(size = 0.5)))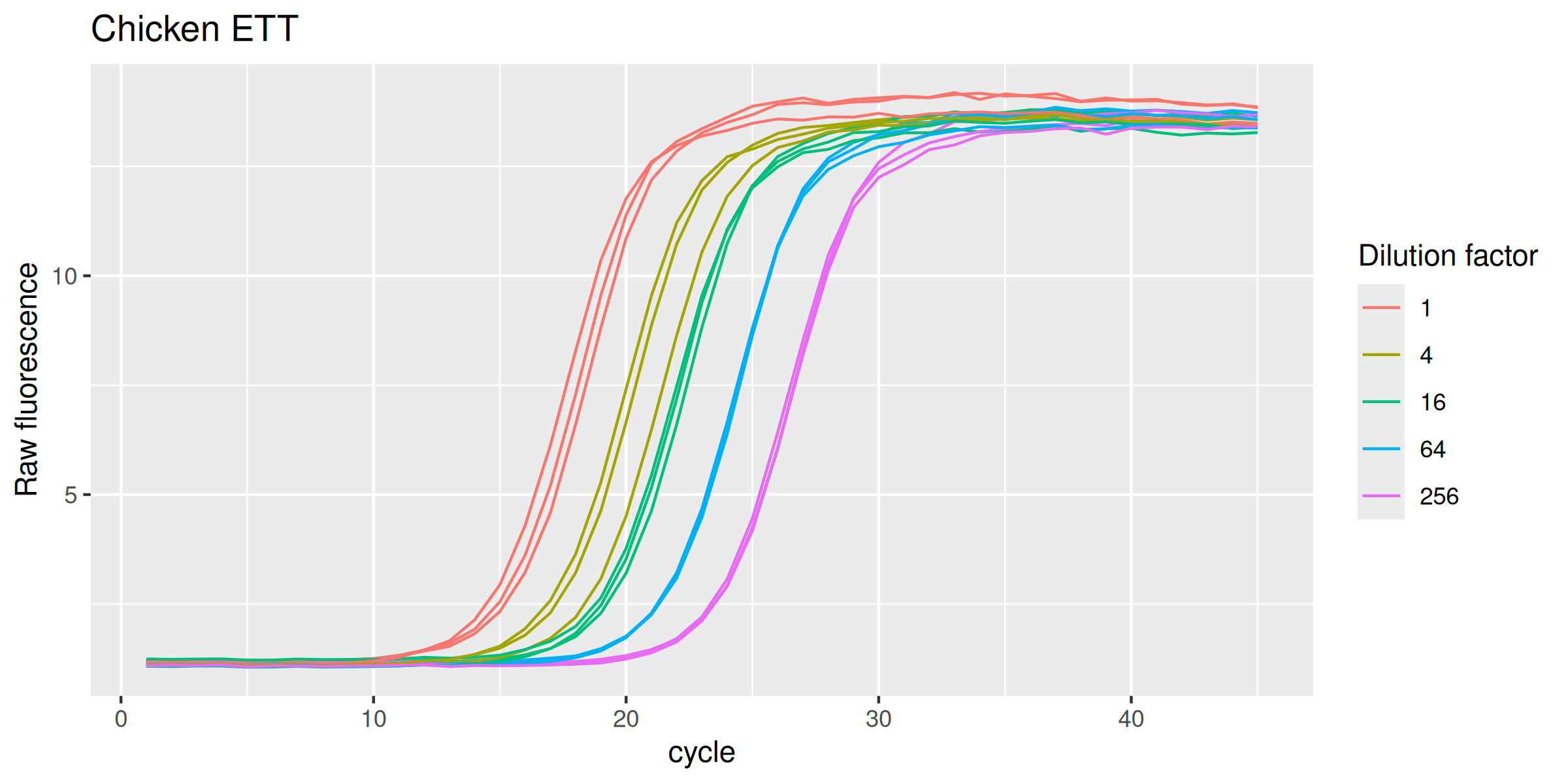
GAPDHh %>%
ggplot(mapping = aes(
x = cycle,
y = fluor,
group = interaction(dilution, replicate),
col = as.factor(dilution)
)) +
geom_line(linewidth = 0.5) +
labs(y = "Raw fluorescence", colour = "Dilution factor", title = "Human GAPDH") +
guides(color = guide_legend(override.aes = list(size = 0.5)))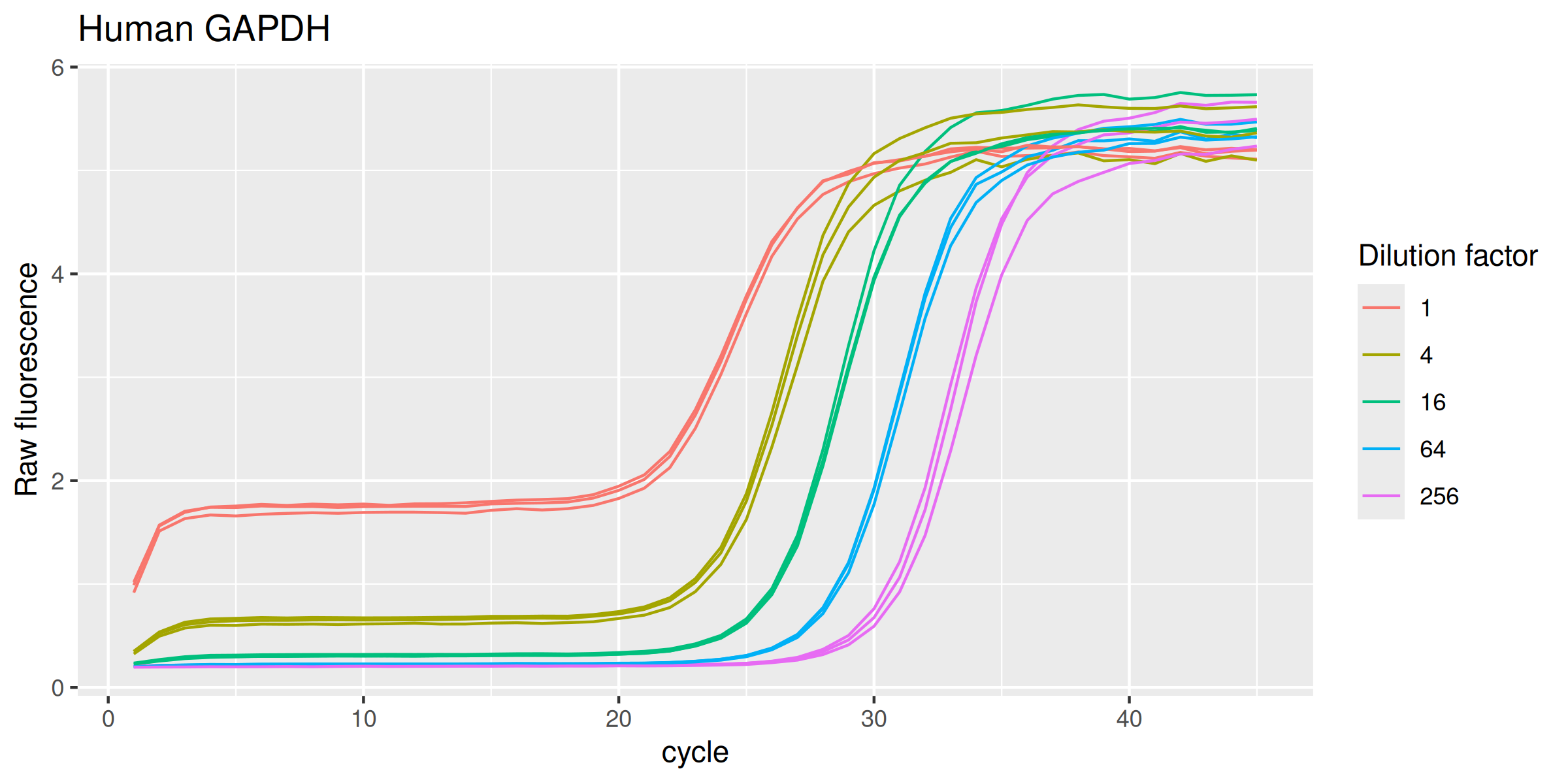
Please note that the {batsch} project is released with a
Contributor
Code of Conduct. By contributing to this project, you agree to abide
by its terms.